While COVID-19 may be transitioning from a “pandemic” to an “endemic” phase, it remains critically important to continue tracking the virus in real-time. In two new papers published in Nature Methods on Feb. 23, 2023, scientists at Scripps Research demonstrate the use of Outbreak.info as a standardized, searchable source of information on the COVID-19 virus and its many variants.
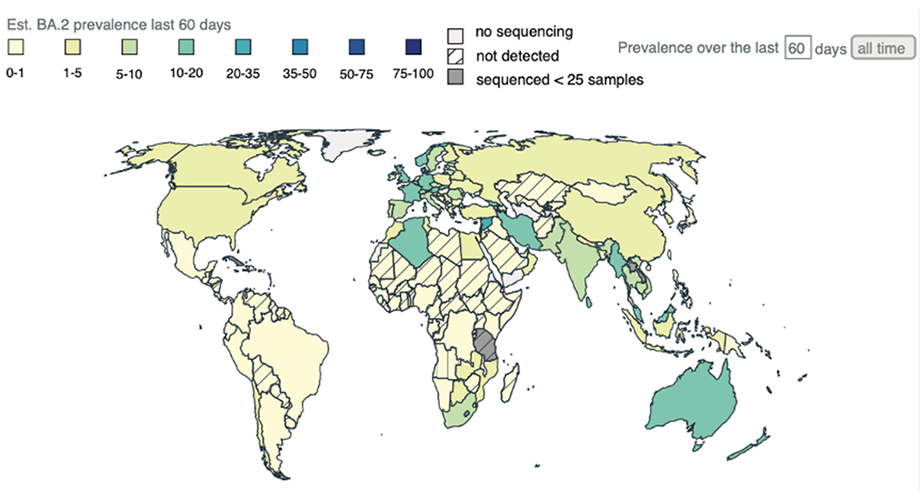
Since the COVID-19 pandemic began, scientists, journalists, public health officials and others interested in tracking the real-time spread and evolution of the coronavirus have used Outbreak.info to access two core features: Daily variant situation reports, which track the prevalence of SARS-CoV-2 variants; and the website’s Research Library, which allows anyone to quickly search through hundreds of thousands of recent and historical COVID-related publications, clinical trials, datasets, protocols and other resources across many sources—regardless of resource type or repository location.
The website was co-developed by multiple Scripps Research scientists, including Institute Investigator Laura Hughes, PhD, Professor Kristian Andersen, PhD, Associate Professor Chunlei Wu, PhD, and Professor Andrew Su, PhD, of the Center for Viral Systems Biology (CViSB). The Center for Viral Systems Biology is a multidisciplinary group of researchers who use genomics, bioinformatics, computational biology, and other areas of research to investigate attributes of viruses that determine the outcomes of disease.
“Since the outbreak of COVID-19 began, the global community—through an unprecedented effort—has sequenced and publicly shared more than 14 million genomes through the GISAID Initiative,” says Hughes. “SARS-CoV-2 variants have dramatically changed the course of the pandemic, with variants like Omicron triggering new waves of cases worldwide. While the unprecedented amount of research on COVID-19 offers new opportunities to track a virus in real-time, analyzing and interpreting such a large volume of data remains a fundamental challenge. Within Outbreak.info, the daily variant reports improve researchers’ ability to follow changes within the SARS-CoV-2 genome and translate them into a better understanding of the virus.”
The Outbreak.info platform currently sifts through data across more than 7,000 locations to track 40 million combinations of Pango lineages, genetically distinct lineages of SARS-CoV-2 based on the analysis of complete or near-complete virus genomes. Importantly, these lineages include SARS-CoV-2 variants of concern, many of which can cause more severe disease.
Additionally, the Research Library aggregates different types of research data into a single searchable platform, allowing researchers to simultaneously discover new publications, clinical trials, datasets and more.
“Many specialized websites have been developed independently to catalog a specific type of COVID-19 research (like publications, for example), but we developed Oubreak.info to be a centralized, standardized repository,” adds Hughes. “Consolidating a wide variety of research all in one place improves researchers’ abilities to discover these resources amidst what has become a massive volume of data, and then translate them into new insights about the virus.”
While the “emergency phase” of the COVID-19 pandemic is winding down, Hughes and Andersen both emphasize that continued research on COVID-19 is essential to understand the virus, track how it is evolving, and improve diagnostics and therapeutics.
“In recent months, many COVID-19 data tracking projects have been forced to scale down or shut down completely as the pandemic has transitioned into an endemic phase,” Andersen says. “However, with lower access to up-to-date information and data, real-time surveillance systems are becoming increasingly more critical to track the virus to understand the emergence and re-emergence of variants. Platforms like Outbreak.info that the global community has come to rely on are essential to support these efforts—not only for SARS-CoV-2, but other emerging pathogens as well.”
In addition to Hughes, Andersen, Wu and Su, Scripps Research authors of the studies, “Outbreak.info Research Library: a standardized, searchable platform to discover and explore COVID-19 resources” and “Outbreak.info genomic reports: scalable and dynamic surveillance of SARS-CoV-2 variants and mutations,” include Karthik Gangavarapu, Alaa Abdel Latif, Julia Mullen, Emory Hufbauer, Ginger Tsueng, Emily Haag, Mark Zeller, Christine M. Aceves, Karina Zaiets, Marco Cano, Jason Lin, Xinghua Zhou, Zhongchao Qian, Rachel Sattler, Nathaniel Matteson, Joshua Levy, Dylan Welzel and Manar Alkuzweny.
Work on Outbreak.info was supported by the National Institute for Allergy and Infectious Diseases, the National Center for Advancing Translational Sciences, the Centers for Disease Control and Prevention and the National Institute of General Medical Sciences.